Targeted Protein Degradation
Multiplexed PROTAC linker optimization with focal Molography
Optimize PROTAC linker design by characterizing 20+ compounds simultaneously on the MACS® Matchmaker. DNA-directed immobilization turns your compound library into a self-assembling biosensor array — delivering structure–activity relationships 20× faster than conventional methods.
The linker optimization bottleneck
PROTACs (proteolysis-targeting chimeras) are transforming drug discovery by enabling selective degradation of disease-relevant proteins via the ubiquitin–proteasome system. The linker connecting the E3 ligase ligand to the protein-of-interest (POI) ligand critically determines binding affinity, ternary complex formation, and drug-like properties.
Systematic optimization of linker structures demands high-throughput affinity characterization across diverse compound series. Traditional methods like SPR or ITC measure one compound at a time, creating a throughput bottleneck that slows the design–make–test–analyze cycle. Chip-to-chip variability in sequential measurements further obscures structure–activity trends.
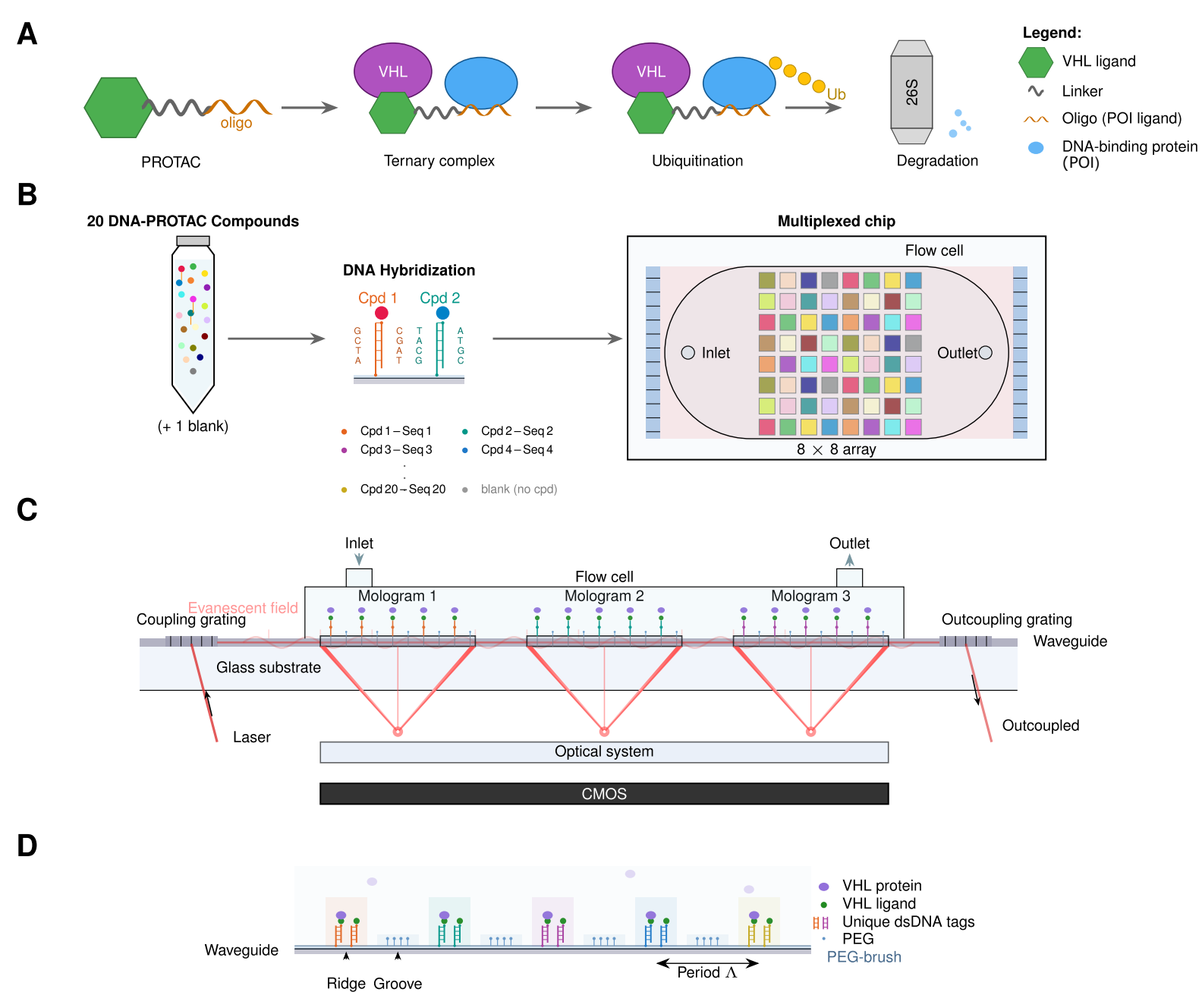
PROTAC mechanism and multiplexed focal Molography workflow. (A) PROTAC-mediated degradation via the ubiquitin–proteasome system. (B) DNA-directed self-assembly of 20 compounds on an 8×8 mologram array. (C) Waveguide-based optical detection. (D) Binding modulation principle.
Multiplexed PROTAC characterization on the MACS® Matchmaker
Focal Molography enables simultaneous measurement of 20–64 DNA-tagged compounds in a single experiment. The DNA-directed immobilization (DDI) workflow eliminates sequential spotting and delivers actionable kinetic data in under 90 minutes.
|
8×8
mologram array
DNA-directed self-assembly
Each compound carries a unique DNA barcode. Watson–Crick base pairing directs compounds to designated positions on the mologram array.
|
5 min
setup time
One-pot immobilization
Pool all compounds and inject as a single mixture. No sequential spotting. No complex array printing.
|
0
non-specific signal
Inherent specificity
Only coherently bound molecules generate signal. Non-specific binding is invisible — no blocking, no reference subtraction.
|
|
20×
throughput increase
Higher throughput
Characterize 20 compounds simultaneously with full kinetic data (KD, kon, koff) in a single run.
|
r = 0.98
method correlation
Stronger SAR signals
Uniform conditions eliminate chip-to-chip variability, yielding dramatically clearer structure–activity trends.
|
Crude
sample matrices
Complex matrices
Measure binding in crude biological samples — serum, cell lysates, tissue extracts — without purification.
|
From compound library to SAR in one experiment
A streamlined four-step protocol turns your DNA-PROTAC library into actionable affinity data.
|
1
SynthesizeBuild DNA-PROTAC library with diverse amino acid linkers via solid-phase chemistry |
2
ImmobilizePool all compounds and inject onto the chip — DDI self-assembles the array in minutes |
3
MeasureFlow target protein at increasing concentrations in single-cycle kinetics format |
4
AnalyzeExtract KD, kon, koff for all compounds simultaneously — ready for SAR |
Accelerate your PROTAC linker optimization
See how multiplexed focal Molography can deliver faster, stronger SAR data for your targeted protein degradation program.
