Profile every Fc receptor on one chip in about one hour.
The Fc Receptor Panel measures FcγRI, FcγRIIa, FcγRIIb, FcγRIIIa, FcγRIIIb, FcεRI, and FcεRII (CD23) in parallel on a single ready-to-use chip, with 8 within-chip replicates per receptor and a built-in oligo-only reference channel for non-specific binding.
7 Fc receptors per chip | 8× Replicates per receptor | ~1h Real-time kinetics across the panel | DDI Pre-conjugated, ready-to-use |
The Fc receptor family and why a single-experiment panel matters.
Activating, inhibitory, and the effector-function profileThe Fcγ receptor family translates antibody binding into cellular effector outputs. FcγRI, FcγRIIa, FcγRIIIa, and FcγRIIIb are activating receptors that drive antibody-dependent cellular cytotoxicity (the lysis of antibody-coated target cells by natural killer cells and macrophages) and antibody-dependent cellular phagocytosis (the engulfment of antibody-opsonized particles). FcγRIIb is the family's only inhibitory receptor and sets the activating-to-inhibitory ratio that underlies most current Fc-engineering campaigns. FcγRIIa carries the H131/R131 polymorphism and FcγRIIIa carries the V158/F158 polymorphism. Both are clinically relevant variants that alter response rates to rituximab, cetuximab, and other approved IgG therapeutics, and comprehensive analytical data on the full FcγR panel is increasingly expected in regulatory submissions for Fc-engineered IgGs and biosimilar comparability under ICH Q5E. | The IgE branch and immune-complex characterizationTwo further Fc receptors are central to allergy biologics, IgE-format diagnostics, and any program with an IgE arm: FcεRI, the high-affinity IgE receptor on mast cells and basophils that triggers allergic effector function, and FcεRII (CD23), a lower-affinity IgE receptor that regulates IgE production and antigen presentation. Both engage the IgE Fc region directly. Anti-IgE biologics, IgE-format diagnostics, and bispecific or fusion formats with an IgE component all need direct FcεRI and CD23 binding data, ideally measured side by side with the same molecule's FcγR profile. This panel pairs the activating and inhibitory FcγRs with FcεRI and CD23 on a single chip, turning what used to be a multi-platform, multi-week characterization into a one-hour multiplex experiment. |
Sequential SPR runs into walls. The Fc panel is built around them.
Modern antibody engineering tunes the entire Fc-receptor profile, not just one receptor at a time, and regulators increasingly expect comprehensive analytical Fc data across the full receptor family. Generating that data on conventional surface plasmon resonance (SPR) means weeks of per-receptor method development and chip-to-chip variability across the panel. Conventional SPR detects refractive-index changes, so specific binding arrives mixed with non-specific adsorption, buffer artifacts, and temperature drift. Focal molography uses a fundamentally different detection geometry: only specifically bound mass that is spatially coherent with the mologram pattern contributes to the readout. Signal and noise separate in Fourier space rather than by subtraction.
⏱ Multi-week SPR → ~1 hour panel Conventional SPR runs each receptor on its own chip on its own day. MACS® Matchmaker carries FcγRI, FcγRIIa, FcγRIIb, FcγRIIIa, FcγRIIIb, FcεRI, and CD23 pre-conjugated on one 8-plex chip, with all seven receptors measured simultaneously in a single antibody injection series. | ∿ Drift-limited → 8 within-chip replicates FcγRII/III and CD23 sit at micromolar KD, where multi-hour SPR drift accumulates over the long association phases needed to fit them. MACS® Matchmaker reports KD, kon, and koff as the mean of 8 independent molograms per receptor, giving confidence intervals from a single experiment and averaging drift out across replicates. |
≈ Non-specific binding (NSB) floor → coherent detection floor Sub-twofold biosimilar comparability and Fc-engineered variant ranking depend on differences smaller than the typical NSB floor of SPR. MACS® Matchmaker reads specifically bound mass by detection geometry, so non-specific binding is incoherent with the mologram pattern and structurally absent from the readout. An 8th oligo-only spot is included on every chip as a built-in reference channel. | ⚙ Method dev. per receptor → pre-conjugated, ready Each SPR receptor needs its own immobilization optimization, regeneration screen, and validation cycle. MACS® Matchmaker ships with all seven Fc receptors plus the oligo reference already DNA-hybridized to the chip via DNA-directed immobilization. No in-house coupling, no per-receptor method development, no regeneration optimization. |
Where the Fc Receptor Panel delivers value.
The same chip and protocol address three distinct workflows across the antibody and immune-complex development process.
Discovery Ranking Fc-engineered candidates Fc engineering tunes the activating-to-inhibitory profile through point mutations in the Fc region. Variants must be ranked across the full receptor family, including the FcγRIIa H131/R131 and FcγRIIIa V158/F158 polymorphisms, and against any IgE-related counterbalance for dual-format biologics. A multiplex panel reveals the whole engineered profile in one experiment per variant. | Biosimilar Comparability Binding equivalence under ICH Q5E Demonstration of FcγR binding equivalence between a biosimilar and its reference is a key element of analytical comparability under ICH Q5E. The affinity differences that must be resolved are often less than twofold, at or below the practical noise floor of conventional surface plasmon resonance. Coherent mass detection plus 8 within-chip replicates is specifically designed for this analytical challenge. | Mechanistic profiling antibodies, immune complexes, IgE biologics For many programs, the relevant question is not how a single Fc receptor binds, but how the whole receptor network responds to the same molecular state: free drug, target-bound, antigen-loaded immune complex. The panel captures this state-dependent receptor footprint and is particularly informative for anti-IgE biologics and bispecific or fusion formats with an IgE component. |
One injection series. Seven receptors. Eight replicates each.
The chip is loaded once with the pre-conjugated Fc-receptor and oligo-only ligand mix via DNA-directed immobilization. Each antibody or immune complex then runs a single ascending-concentration injection series in which all seven Fc receptors and the oligo reference are read simultaneously. The running buffer wash regenerates the surface between samples, so no separate regeneration step is required.
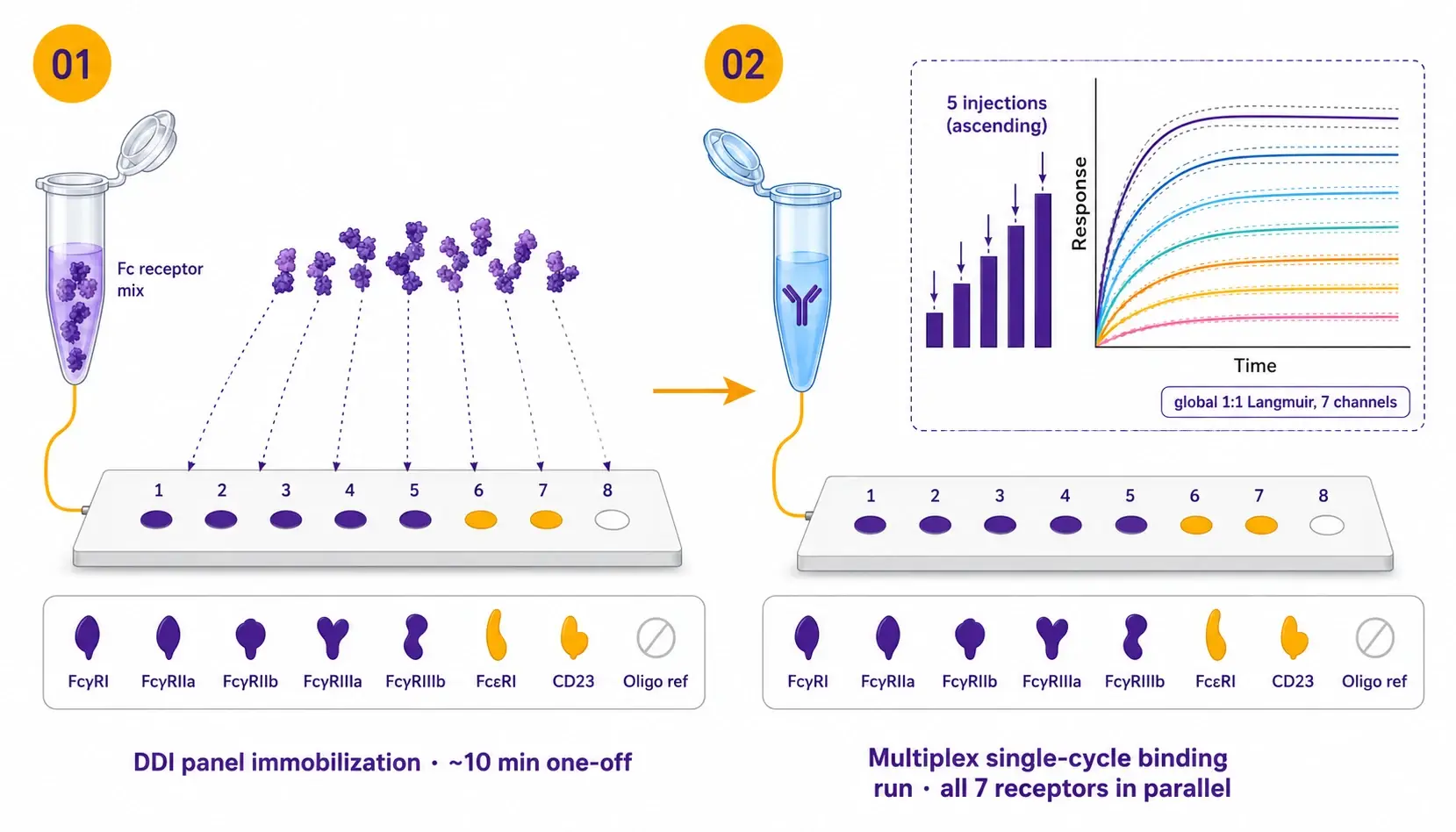 | Protocol details
|
Seven receptors. Eight replicates. One hour.
See the Fc Receptor Panel on a real antibody. Your antibody. Book a 30-minute demo with one of our application scientists.
What makes MACS® Matchmaker the purpose-built solution for this workflow.
7 receptors The full Fc receptor family in one run FcγRI, FcγRIIa, FcγRIIb, FcγRIIIa, FcγRIIIb, FcεRI, and FcεRII (CD23) profiled under identical assay conditions in the same injection series, eliminating the inter-assay variability of sequential chip runs. | 8 replicates Within-chip statistics from a single experiment Reported KD, kon, and koff are the mean of 8 independent molograms per receptor, with confidence intervals derived from global fitting across all channels. | ~1 hour Real-time kinetics across the full panel Single-cycle ascending-concentration kinetics on the multiplex chip. No regeneration step between antibodies; the surface clears with the running buffer. |
No prep Chips ship pre-conjugated and ready to use All seven Fc receptors plus an oligo-only reference are immobilized via DNA-directed immobilization on the chip prior to shipment. No in-house coupling, no surface optimization, no batch-to-batch ligand variability. | pg/mm² A direct measurement of specifically bound mass Coherent mass density is not a proxy for binding. It is a direct physical readout of the mass that binds coherently to the mologram pattern, feeding directly into 1:1 Langmuir kinetic analysis without refractive-index conversion. | Built-in NSB ref Every chip is self-validating An 8th oligo-only construct with no captured Fc receptor occupies one position on every chip. Authentic Fc-mediated signals on positions 1 to 7 should produce no response on this channel: a built-in non-specific-binding integrity check on every run. |
MACS® Matchmaker vs. competing HT-SPR platforms.
High-throughput SPR platforms excel at screening many antibodies against one receptor, a complementary application. The comparison below addresses the multi-receptor Fc-panel workflow specifically.
| Parameter | MACS® Matchmaker | Biacore SPR / Octet BLI / Carterra HT-SPR |
|---|---|---|
| Full Fc panel (FcγR + IgE arm) | ✓ All 7 receptors in parallel, ~1 hour, 1 chip | Sequential, multiple days, multiple chips |
| Replicates per receptor | ✓ 8 within-chip molograms | Not built-in; requires repeat runs |
| Non-specific binding | ✓ Structurally absent (Fourier-space) + 8th oligo reference | Subtracted via reference channel |
| Signal readout | ✓ Coherent mass density (pg/mm²) | Refractive index units (proxy) |
| Surface preparation | ✓ Pre-conjugated, ready to use | In-house ligand arraying required |
| Allotypes (H131/R131, V158/F158) | ✓ Integrated as separate panel positions | Each variant requires its own chip |
| Sub-twofold ICH Q5E resolution | ✓ Coherent detection floor | Constrained by NSB residual |
See the panel resolve a Fab-to-Fc allosteric coupling on a real biologic.
A peer-quality application note profiling four states of the anti-IgE biologic omalizumab on the Fc Receptor Panel: free omalizumab, free IgE, the binary IgE/omalizumab complex, and the ternary IgE/omalizumab/allergen complex. Hero finding: a residual 292 pM FcεRI signal on the binary complex that switches off when the allergen loads the complex, while every FcγR rescales. Complete dataset, fits, methods, and citable references.
Request the AppNote →