Predict human antibody half-life across all six preclinical species in one hour.
The FcRn Species Panel delivers kinetic and equilibrium constants for human, cynomolgus, marmoset, mouse, rat, and minipig FcRn simultaneously, on a single ready-to-use chip, in approximately one hour.
FcRn-mediated antibody recycling and the challenge of cross-species pharmacokinetics.
The role of FcRn in antibody pharmacokineticsThe neonatal Fc receptor (FcRn) governs the serum half-life of immunoglobulin G (IgG) through a pH-dependent recycling mechanism. Following pinocytosis, IgG molecules are internalized into acidified endosomes (pH ~6.0), where FcRn binds with high affinity and rescues the antibody from lysosomal degradation. Upon fusion with the cell membrane at physiological pH (~7.4), the interaction reverses and intact IgG is returned to circulation. 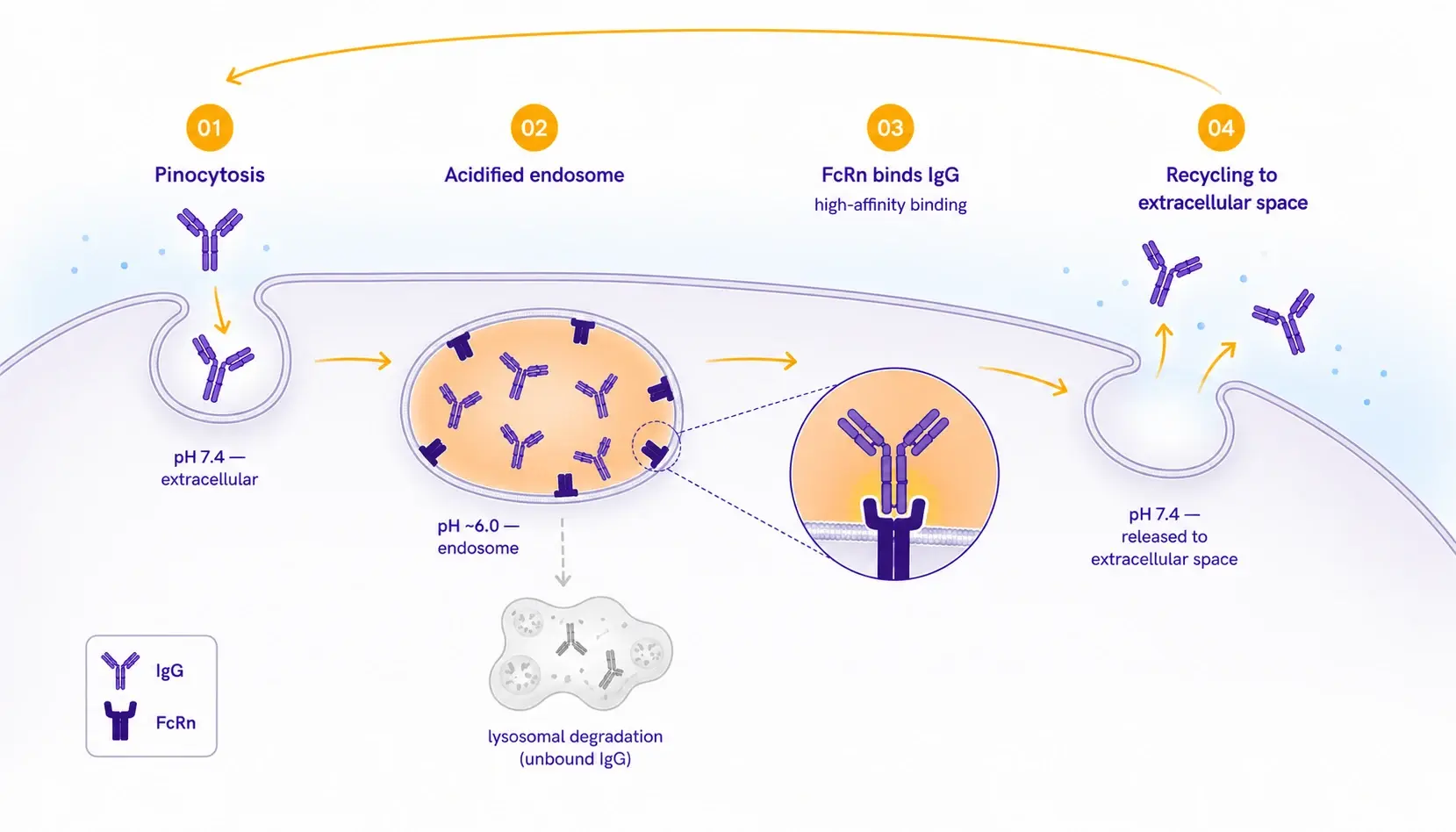 For therapeutic antibody developers, FcRn affinity at pH 6.0 is a critical developability attribute, directly linked to dosing frequency, patient exposure, and the success of half-life-extending Fc engineering strategies, which introduce point mutations into the Fc region to tighten FcRn binding. Characterizing this interaction accurately and early in development directly informs lead selection and preclinical study design. | Why species selection matters for PK predictionA central challenge in translational pharmacokinetics is that FcRn affinity differs substantially across preclinical species. The mouse-human gap can reach 5- to 10-fold for the same antibody, and rat FcRn binding is similarly attenuated. Together, these rodent-human discrepancies account for the systematic under-prediction of human half-life from rodent pharmacokinetic (PK) studies. Allometric PK scaling, the standard practice of predicting human pharmacokinetics from animal data through body-size relationships, presumes consistent receptor biology across species. If the FcRn affinity of your molecule in the toxicology species differs substantially from its affinity in human, that presumption breaks and the human PK projection becomes unreliable. |
SPR runs into walls. MACS® Matchmaker is built around them.
FcRn affinity varies by 5–10× between mouse and human for the same antibody, and by as little as 1.5× between cynomolgus and human. The larger gap drives systematic under-prediction of human half-life from rodent studies; the smaller one, alongside ICH Q5E biosimilar comparability (the international guideline on analytical similarity for biotherapeutics), lives below the residual noise floor of conventional SPR. Conventional SPR detects refractive-index changes, so specific binding arrives mixed with non-specific adsorption (NSB), temperature drift, and buffer-composition shifts. Focal molography reads specifically bound mass directly by spatial coherence, separating signal and noise in Fourier space rather than by subtraction.
⏱ Multi-week SPR → ~1 hour panel Conventional SPR runs each species on its own chip on its own day. MACS® Matchmaker carries human, cyno, marmoset, mouse, rat, and minipig FcRn pre-conjugated on one 8-plex chip, with all six species measured simultaneously in a single antibody injection series. | ∿ Drift-limited → 8 within-chip replicates FcRn KD values sit at 0.5–10 µM; multi-hour SPR drift accumulates over the long association phases needed to fit them. MACS® Matchmaker reports KD, kon, koff as the mean of 8 independent molograms per species, giving confidence intervals from a single experiment and averaging drift out across replicates. |
≈ NSB floor → coherent detection floor Sub-twofold biosimilar comparability and cross-species ratio analysis depend on differences smaller than the typical NSB floor of SPR. MACS® Matchmaker reads specifically bound mass by detection geometry, so non-specific binding is incoherent with the mologram pattern and structurally absent from the readout. | ⚙ Method dev. per receptor → pre-conjugated, ready Each SPR species needs its own immobilization optimization, regeneration screen, and validation cycle. MACS® Matchmaker ships with all eight ligands already DNA-hybridized to the chip; the pH 7.4 wash dissociates analyte completely, so no regeneration step or per-receptor method development is required. |
Where the FcRn Species Panel delivers value.
The same chip and protocol address three distinct workflows across the therapeutic antibody development process.
Discovery Ranking Fc-engineered candidates Fc engineering for half-life extension requires ranking variants not only on human FcRn but across the preclinical species intended for PK and toxicology studies. Affinity gains observed on human FcRn are not always preserved to the same degree in mouse and rat. Identifying these discrepancies before in vivo studies prevents costly course-corrections. | Developability Cross-species PK bridging and allometric scaling FcRn affinity differences of 5- to 10-fold between mouse and human account for systematic under-prediction of human antibody half-life from murine PK studies. Characterizing the complete cross-species affinity profile early in development provides the data needed to calibrate allometric PK models and to select the most predictive tox species. | Biosimilar Comparability Binding equivalence under ICH Q5E Demonstration of FcRn binding equivalence between a biosimilar and its reference is a key element of analytical comparability under ICH Q5E. The affinity differences that must be resolved are often less than twofold, at or below the practical noise floor of conventional SPR. Coherent mass detection plus 8 within-chip replicates is specifically designed for this analytical challenge. |
The pH-switch single-cycle kinetics protocol.
The chip is loaded once with the pre-conjugated FcRn–oligo ligand mix via DNA-directed immobilization. Each antibody then runs a pH 7.4 specificity check followed by an ascending-concentration pH 6.0 binding run on the same chip. The pH 7.4 wash regenerates between antibodies, so no separate regeneration step is required.
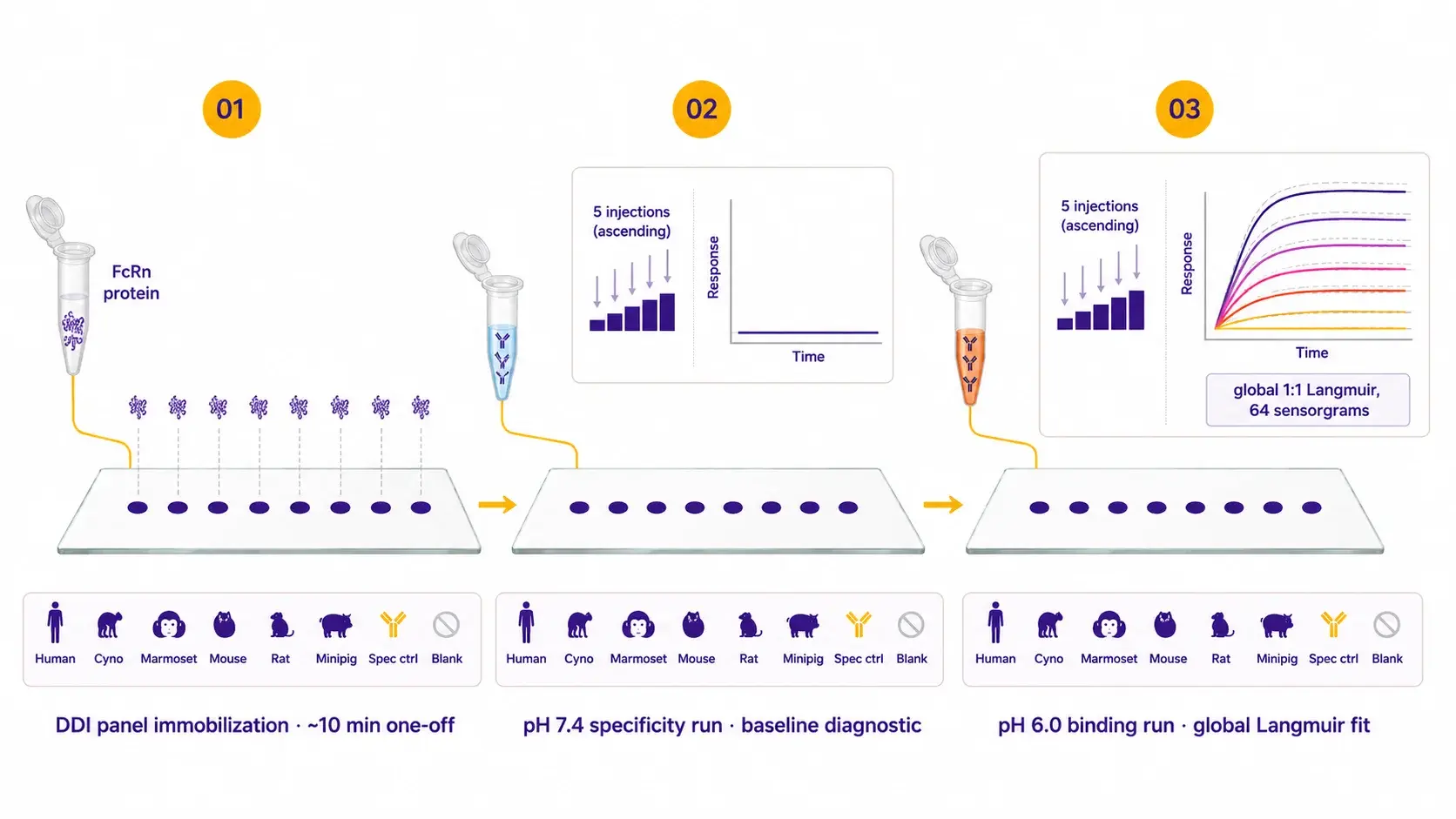 | Protocol details
|
Six species. Eight replicates. One hour.
See the FcRn Species Panel on a real antibody. Your antibody. Book a 30-minute demo with one of our application scientists.
MACS® Matchmaker vs. competing HT-SPR platforms.
High-throughput SPR platforms excel at screening many antibodies against one ligand: a complementary application. The comparison below addresses the cross-species FcRn workflow specifically.
| Parameter | MACS® Matchmaker | Biacore SPR / Octet BLI / Carterra HT-SPR |
|---|---|---|
| Cross-species profiling | ✓ All 6 species in parallel, ~1 hour, 1 chip | Sequential, multiple days, multiple chips |
| Replicates per species | ✓ 8 within-chip molograms | Not built-in; requires repeat runs |
| Non-specific binding | ✓ Structurally absent (Fourier-space) | Subtracted via reference channel |
| Signal readout | ✓ Coherent mass density (pg/mm²) | Refractive index units (proxy) |
| Surface preparation | ✓ Pre-conjugated, ready to use | In-house ligand arraying required |
| Built-in specificity control | ✓ IgG-binding-deficient FcRn on every chip | None standard |
| Sub-twofold ICH Q5E resolution | ✓ Coherent detection floor | Constrained by NSB residual |
What makes MACS® Matchmaker the purpose-built solution for this workflow.
6 species All relevant preclinical species in one run Human, cynomolgus, marmoset, mouse, rat, and minipig FcRn profiled under identical assay conditions in the same injection series, eliminating the inter-assay variability of sequential chip runs. | 8 replicates Within-chip statistics from a single experiment Reported KD, kon, and koff are the mean of 8 independent molograms per species, with confidence intervals derived from global fitting of all 64 sensorgrams simultaneously. | ~1 hour Binding kinetics and specificity in a single session The pH 6.0 binding run and pH 7.4 specificity run are performed sequentially on the same chip, with no regeneration step required between them. |
No prep Chips ship pre-conjugated and ready to use All six FcRn species are immobilized via DDI on the Oligo|PEG surface prior to shipment. No in-house coupling chemistry, no surface optimization, no batch-to-batch ligand variability. | pg/mm² A direct measurement of specifically bound mass Coherent mass density is not a proxy for binding. It is a direct physical readout of the mass that binds coherently to the mologram pattern, feeding directly into 1:1 Langmuir kinetic analysis without refractive-index conversion. | Built-in QC Every chip is self-validating An IgG-binding-deficient FcRn variant occupies one position on every chip. Authentic FcRn-mediated signals on positions 1–6 should produce no response on this channel: a built-in assay integrity check on every run. |
Get the complete cross-species reference dataset.
A peer-quality application note compiling cross-species FcRn KD values, half-life-extended Fc variant rankings (YTE, LS), full methods, sensorgram library, and a citable bibliography.
Request the AppNote →