Catch fast clearance, aggregation, and immunogenicity before they kill your drug candidate.
The Polyreactivity Panel quantifies non-specific binding across 8 or 16 defined liability axes in a single label-free injection on the same MACS® Matchmaker chip and sample as your target binding kinetics. Score every candidate against the Jain et al. 2017 clinical-stage mAb benchmark.
8 or 16 Liability axes per chip | ~30 min Single SCK injection | 137 mAbs Jain clinical-stage benchmark | DDI Pre-conjugated, ready-to-use |
Polyreactivity is a top developability liability, and it is currently measured off-instrument.
Why polyreactivity drives clinical attritionPolyreactive antibodies, those that bind structurally unrelated antigens through non-specific surface chemistry, exhibit accelerated serum clearance, elevated immunogenicity risk, self-association, and aggregation. Hötzel and colleagues showed that even modest polyreactivity correlates with poor pharmacokinetics in human subjects, and Jain et al.'s 2017 survey of 137 clinical-stage monoclonal antibodies established that polyreactivity scores correlate with progression through clinical development. The biophysical liability is set by surface charge patches, hydrophobic residues, and non-specific affinity for membranes, nucleic acids, and abundant plasma proteins. Once a polyreactive lead enters lead optimization, the cost of catching it later, in pharmacokinetic (PK) studies, in toxicology, in early clinical, can be measured in months and millions. | Industry-standard methods are off-instrument and slowStandard polyreactivity readouts are plate-based: PSR-ELISA on a soluble-membrane preparation, AC-SINS on aggregated antibody, BV-ELISA on baculovirus particles. Each consumes tens of micrograms of candidate per measurement, takes hours per plate, and runs on a different instrument than the target binding assay. The result is that polyreactivity is measured after affinity ranking is complete, often after a lead candidate has been committed. The MACS® Matchmaker Polyreactivity Panel runs on the same chip and the same sample as your target binding kinetics, with the same fluidics and the same data export, in approximately 30 minutes per candidate. |
Plate-based polyreactivity assays break the discovery cycle. The panel is built to fit it.
Polyreactivity has to be screened during candidate selection, not after. Conventional methods are too slow and too material-hungry to be run on every candidate at every cycle, so polyreactivity is typically measured only on a short-list, late, when course corrections are expensive. The Polyreactivity Panel uses the same focal molography surface as the target-binding assay: defined ligands immobilized via DNA-directed immobilization (DDI), a single NHS-amine conjugation chemistry, and the same single-cycle kinetics protocol. The polyreactivity readout drops into the same pipeline, the same buffer, the same sample volume, the same instrument.
⏱ Off-instrument plate methods → 30-min on-instrument PSR-ELISA, AC-SINS, and BV-ELISA each take hours per plate and run separately from the target-binding assay. MACS® Matchmaker runs the polyreactivity panel as a single SCK injection on the same chip as the affinity assay, in approximately 30 minutes per candidate. | ⤆ Material-hungry → micrograms Plate methods consume tens of micrograms of antibody per candidate per readout. MACS® Matchmaker runs the polyreactivity panel on the same low-volume sample injection used for target binding, freeing material for the kinetic assay and downstream characterization. |
≈ Heterogeneous reagents → defined ligands Soluble-membrane PSR preparations and aggregated antibody preps are heterogeneous and lot-to-lot variable. MACS® Matchmaker uses commercial, recombinant or highly purified ligands plus defined amine-functionalised lipids and glycosaminoglycans, all conjugated through a single NHS chemistry. No aggregates, no membrane preparations, no carrier-protein conjugates. | ⚙ Late-stage flag → at every discovery cycle Off-instrument workflows force polyreactivity to be measured only on a triaged short-list. MACS® Matchmaker lets the polyreactivity panel run on every candidate at every cycle, alongside affinity, surfacing developability liabilities at the earliest decision point. |
Where the Polyreactivity Panel delivers value.
The same chip, sample, and protocol address three distinct workflows across antibody discovery and developability.
Discovery Drop polyreactive candidates before lead optimization Run the panel on every candidate alongside the target affinity measurement. Surface polyreactivity at the earliest selection point, before resources are committed to lead optimization. The Polyreactivity Index converts the multi-axis readout into a single quantitative number that can be paired directly with KD in candidate-ranking tables. | Lead Optimization Rank engineered variants on the developability axis Fc engineering, charge-patch removal, and CDR optimization can each shift the polyreactivity profile in unintended directions. The panel ranks engineered variants on the polyreactivity axis with the same within-chip statistics as the affinity measurement, in the same experiment. | Developability Score every candidate before lead-nomination Build the Polyreactivity Index into the standard developability data package alongside FcRn, target KD, hydrophobic interaction chromatography, and self-interaction. Anchored to the Jain 2017 clinical-stage benchmark distribution, the PI gives a calibrated risk score that travels with the candidate into formulation and IND-enabling work. |
One workflow. Two depths of liability coverage.
Polyreactivity Essentials, 8-plexThe triage configuration. One ligand per major liability axis: Protein A/G as the Fc-capture positive control, dsDNA reference, human serum albumin, insulin, lysozyme, aminocardiolipin, aminoheparin, and polyclonal human IgG. 8 within-chip replicates per ligand for high-confidence statistics from a single experiment. Built for the routine candidate-triage workflow at every discovery cycle. Drop polyreactive candidates early without consuming the material and the time of a full developability panel. | Polyreactivity Plus, 16-plexThe in-depth configuration. Adds parallel ligands per axis (BSA + HSA, ovalbumin + insulin, ssDNA + dsDNA), the antiphospholipid-syndrome autoantigen β2-glycoprotein I, lipopolysaccharide, transferrin, human IgM, and fibrinogen. 4 within-chip replicates per ligand, with the breadth needed for cross-axis liability mapping. Built for the developability assessment that supports lead-nomination and IND-enabling work, where the breadth of axes resolves which liability is driving the score and informs targeted engineering. |
One injection. All axes. Approximately 30 minutes.
The chip is loaded once with the pre-conjugated ligand panel via DNA-directed immobilization. Each candidate then runs a single ascending-concentration injection series in HBS-EP+ pH 7.4, with the polyreactivity response read simultaneously across all 8 or 16 channels. The running buffer wash regenerates between candidates for routine screening; a brief glycine pH 2.0 strip recovers the Protein A/G channel for high-throughput cycles.
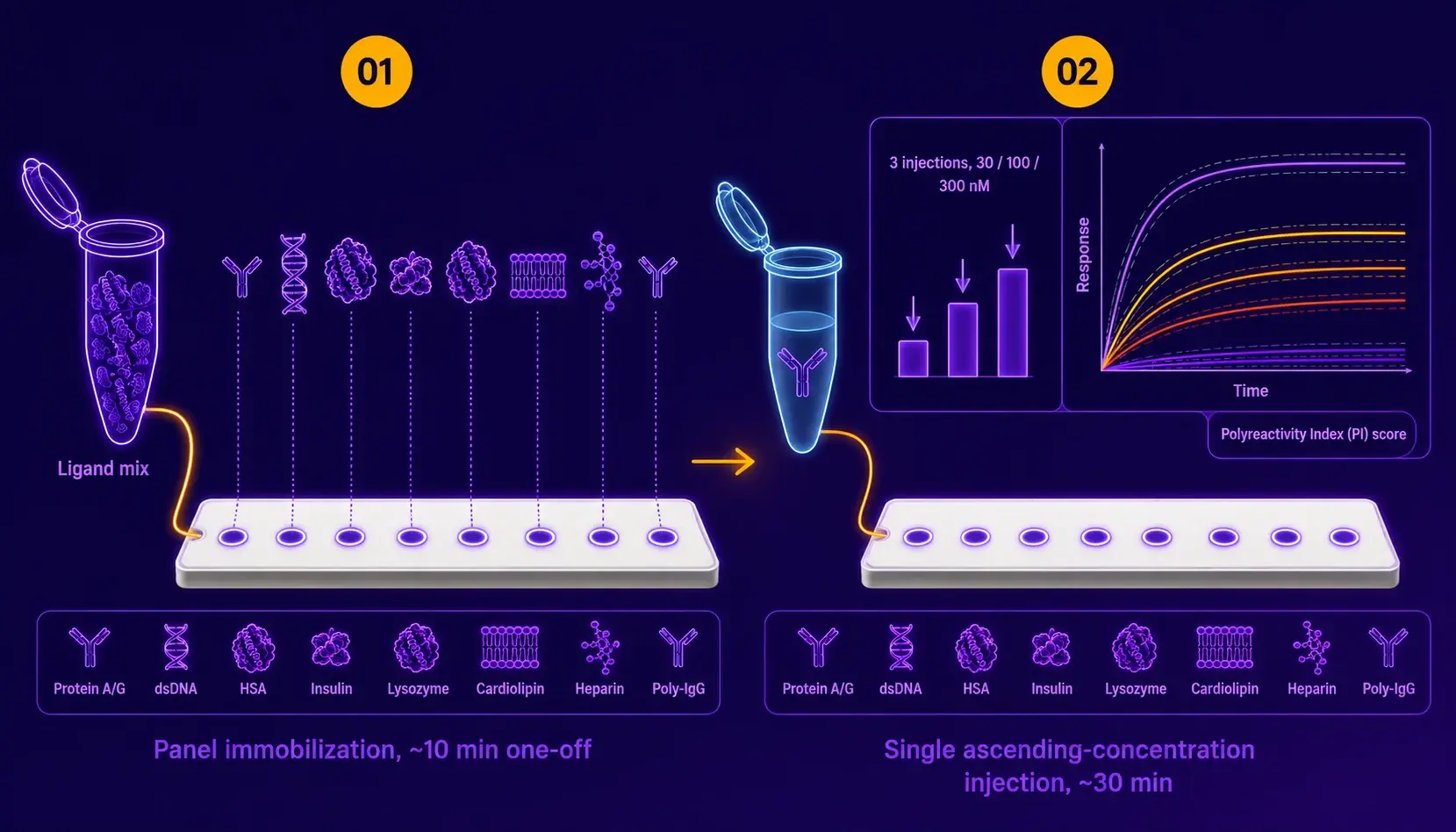 | Protocol details
|
Eight axes or sixteen. Thirty minutes either way.
Run the Polyreactivity Panel on a real candidate. Your candidate. Book a 30-minute demo with one of our application scientists.
What makes MACS® Matchmaker the purpose-built solution for this workflow.
8 or 16 axes Two configurations for two use cases Polyreactivity Essentials (8 ligands, one per major liability axis) for routine triage; Polyreactivity Plus (16 ligands, parallel axes plus β2-GPI, LPS, IgM, fibrinogen) for in-depth developability. Both run on the same instrument with the same protocol. | Quantitative PI Anchored to clinical-stage benchmarks The Polyreactivity Index is the mean equilibrium response across non-control channels, normalized to the Protein A/G control. Three-tier thresholds (acceptable / orthogonal-confirm / liability) are anchored to the Jain et al. 2017 distribution of 137 clinical-stage monoclonal antibodies. | Single chemistry NHS-conjugated DDI All ligands are conjugated through a single NHS amine-reactive chemistry to amine-modified DDI oligos. No aggregates, no soluble-membrane preparations, no carrier-protein hapten conjugates. Lot-to-lot reproducibility designed in. |
~30 min Same chip as target binding The polyreactivity panel runs on the same MACS® Matchmaker chip and the same sample injection used for the target affinity assay. Polyreactivity moves into the discovery cycle instead of running parallel to it. | pg/mm² Direct mass detection Coherent mass density at each ligand spot is a direct physical readout of bound mass, not a refractive-index proxy. The Polyreactivity Index is therefore a quantitative number rather than a relative semi-quantitative score. | Built-in QC Protein A/G plus replicates The Protein A/G channel provides a per-injection Fc-capture quality control: absence of signal flags candidate or run failure before any PI is computed. 8 (Essentials) or 4 (Plus) within-chip replicates per ligand provide confidence intervals from a single experiment. |
MACS® Matchmaker vs. plate-based polyreactivity methods.
PSR-ELISA, AC-SINS, and BV-ELISA are the established plate methods. The comparison below addresses the developability-screening workflow specifically.
| Parameter | MACS® Matchmaker | PSR-ELISA / AC-SINS / BV-ELISA |
|---|---|---|
| Time per candidate | ✓ ~30 min single chip | hours of plate work |
| Sample consumption | ✓ µg-scale, shared with target assay | 10s of µg per candidate per readout |
| Same workflow as target binding | ✓ Yes, same chip, same sample | No, separate instrument and prep |
| Liability axes covered | ✓ 8 or 16, defined ligands | 1 to 3 per assay (axis-specific) |
| Reagent definition | ✓ Recombinant or purified, single NHS chemistry | Soluble-membrane preps, aggregated antibody, baculovirus particles |
| Replicates per ligand | ✓ 8 (Essentials) / 4 (Plus) within-chip | Plate triplicate typical |
| Quantitative score | ✓ Polyreactivity Index, pg/mm² readout | Semi-quantitative or relative |
Get the full panel composition and threshold validation.
A peer-quality application note describing the two-configuration panel design, the per-axis ligand rationale, the Polyreactivity Index calculation, and the threshold anchoring against the Jain 2017 clinical-stage mAb distribution. Includes the trastuzumab / adalimumab low-PI controls and the 4E10 / 2F5 HIV broadly neutralizing antibody high-PI controls used to validate the assay.
Request the AppNote →