Optimize PROTAC linkers 20× faster, in one experiment.
Characterize 20 to 64 DNA-tagged PROTAC (proteolysis-targeting chimera) variants and molecular glues simultaneously on a single chip. Reveal structure-activity relationships across linker chemistry in 90 minutes, with kinetics readable directly in cell lysate.
Targeted protein degradation and the linker SAR bottleneck.
How PROTACs and molecular glues workPROTACs (proteolysis-targeting chimeras) and molecular glues are heterobifunctional or monovalent small molecules that recruit an E3 ubiquitin ligase to a protein of interest (POI). The resulting ternary complex (POI · degrader · E3) is ubiquitinated and shuttled to the 26S proteasome for degradation. Unlike inhibitors, degraders act catalytically and have access to scaffolding proteins, transcription factors, and other historically undruggable targets. 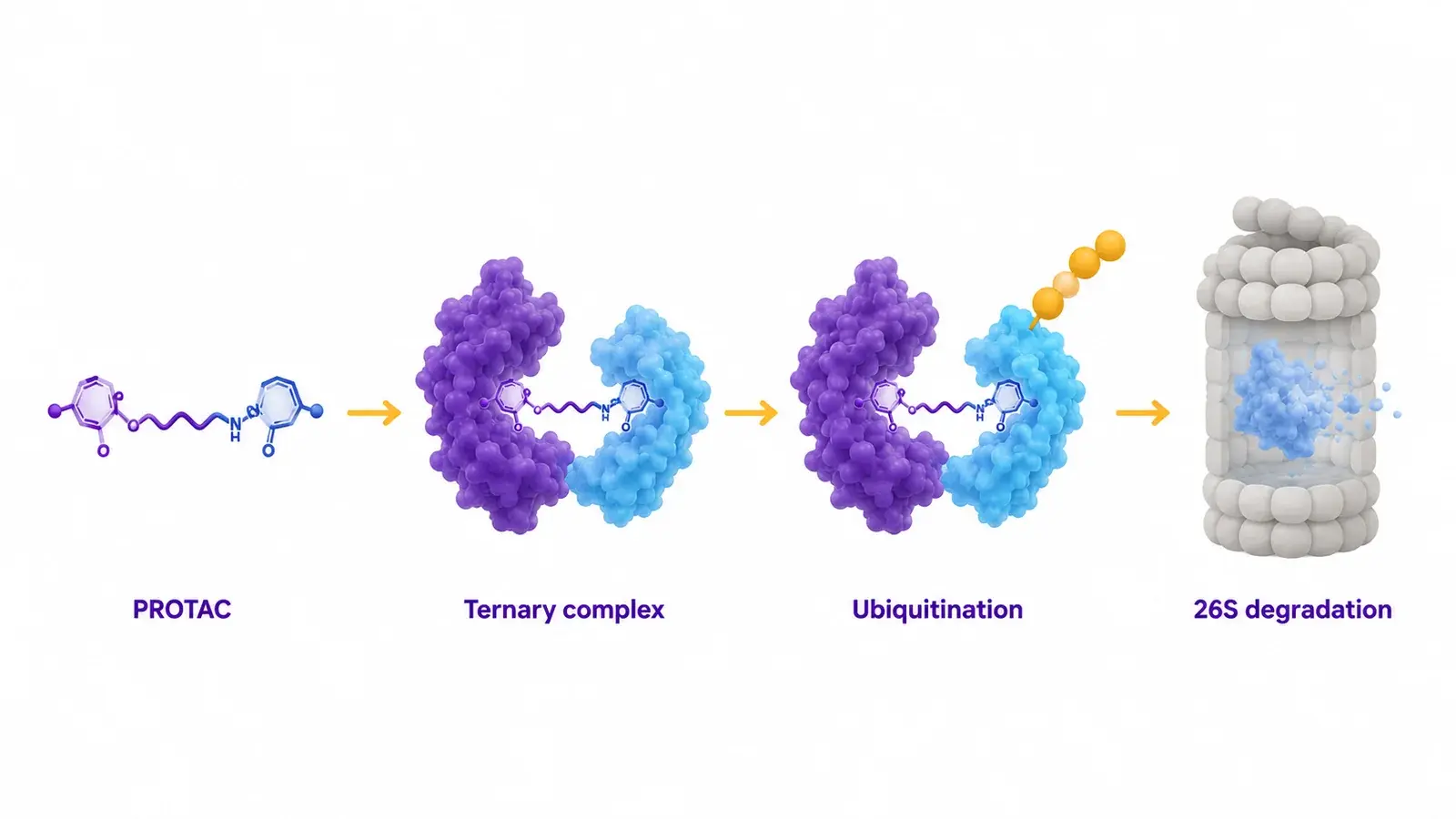 The linker that bridges the E3 ligand and the POI ligand is the central design parameter. Length, rigidity, and lipophilicity govern ternary complex geometry, cooperativity (α), and ultimately the cellular degradation efficiency. Small linker changes routinely produce 10-fold to 100-fold shifts in DC50, so systematic linker SAR is non-negotiable in lead optimization. | Why linker SAR is hard to measureConventional biophysics characterizes one compound per chip per day. A 20-compound linker series therefore consumes a full week of instrument time, with chip-to-chip drift adding variance that obscures the underlying SAR. Buffer-only KD values frequently fail to predict cellular degradation because cooperativity is matrix-dependent and many E3 ligases lose stability after purification. Cellular degradation assays read the integrated outcome but provide no biophysical interpretation, leaving designers without kinetic feedback on which step (binding, cooperativity, geometry) drove the change. The result is iterative empirical screening when a single multiplexed kinetic measurement could resolve the design space in one run. |
One chip. 20 linkers. Lysate-compatible.
Linker SAR demands many compounds, real cooperativity, and the freedom to use E3 ligases in their native context. MACS® Matchmaker addresses each constraint in a single experiment by combining DNA-directed immobilization (DDI) with coherent mass detection.
⏱ Sequential SPR weeks → 90-minute panel Conventional SPR runs each linker variant on its own chip on its own day. MACS® Matchmaker carries 20 to 64 DNA-tagged compounds on one chip, all measured under identical conditions in a single concentration series. | ∿ Chip-to-chip drift → within-chip uniformity Multiplex measurement on one chip removes the largest source of SAR noise. The Würzburg DNA-VHL study reported method correlation of r = 0.98 across multiplexed equilibrium and kinetic readouts, versus 0.88 for the matched singleplex experiment. |
≈ Buffer-only KD → lysate-native ternary Coherent mass detection rejects non-specific binding by detection geometry, so E3 ligases can be presented in cell lysate where many remain stable. The recombinant-only constraint of conventional SPR no longer limits assay design. | ⚙ Sequential spotting → one-pot loading All compounds are pooled and injected once. Watson-Crick hybridization directs each barcode to its capture position within five minutes. No printing, no sequential coupling, no per-compound regeneration. |
Where multiplexed focal Molography accelerates your degrader program.
The same chip and protocol address three workflows that span the full TPD discovery pipeline.
Linker SAR optimization A 20-compound linker series characterized in 90 minutes, with kinetic constants reported per compound and physicochemical descriptors directly correlatable with binding. Identify the linker design rule (lipophilicity, length, rigidity) that drives your specific E3 / POI pair, in one experiment. | Molecular glue hit validation Glue-induced ternary interactions are cooperative and matrix-dependent, often invisible in clean buffer. The drift-free baseline and lysate compatibility resolve cooperativity values (α) and slow off-rates that classical equilibrium binding misses, including 14-3-3 PPI glues and non-degrading RapaGlue-type binders. | DEL hit-to-lead for degraders DNA-encoded library hits arrive on-DNA. The same DNA tag that identified the hit also drives immobilization on the mologram. Hits move from selection to validated kinetic profile without off-DNA resynthesis, with E3 or POI as the analyte. |
From DNA-tagged compound library to ranked SAR.
The 8×8 mologram array is loaded once via DDI of the pooled compound mix. The target protein (or lysate containing it) is then injected at five ascending concentrations in a single-cycle kinetics format. Both equilibrium and kinetic KD are extracted in the same run, and physicochemical descriptors (cLogP, cLogS, polar surface area) are correlated with binding directly.
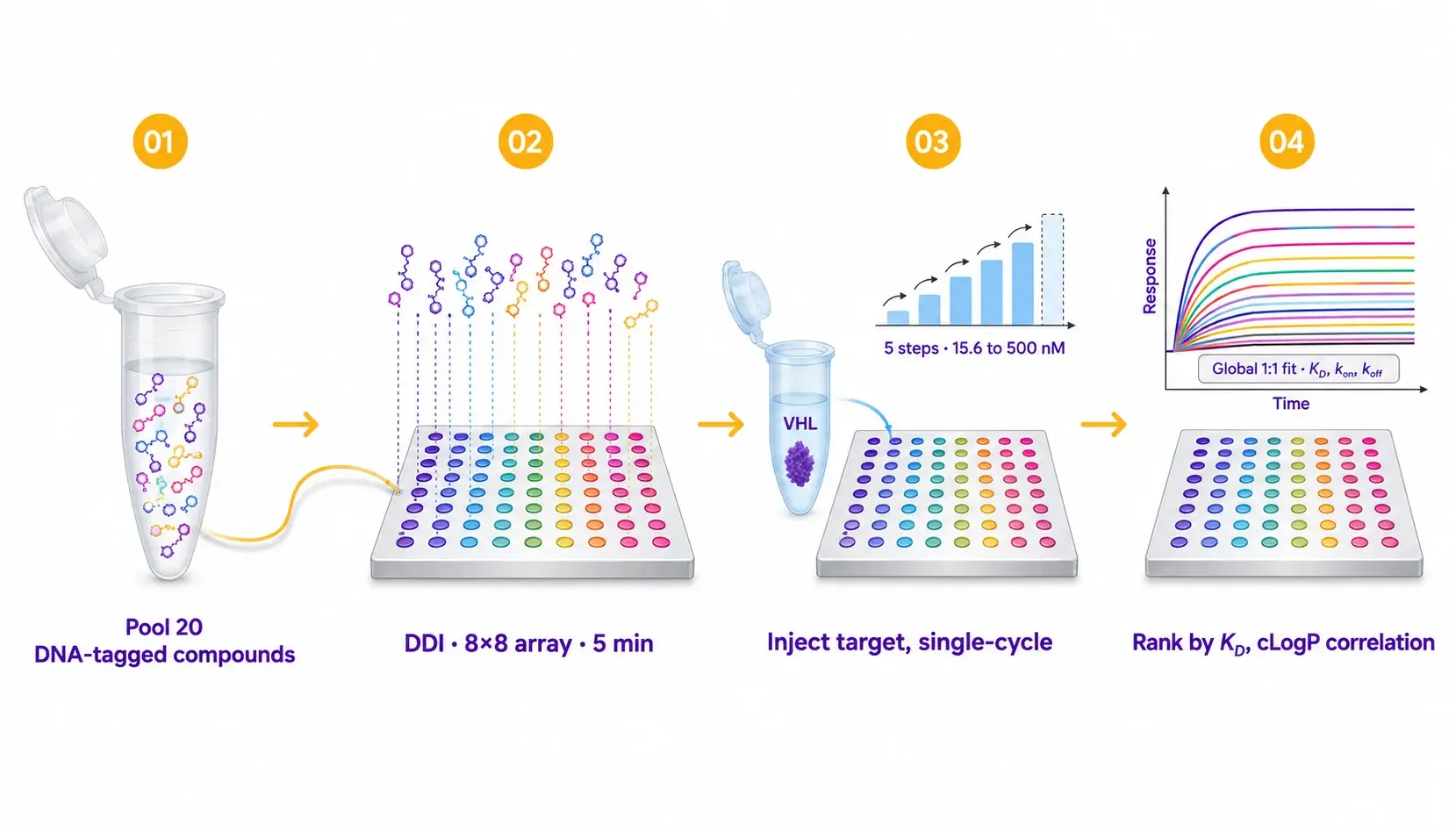 | Protocol details
|
20 linkers. One chip. 90 minutes.
Send us your DNA-tagged PROTAC library and your target. Get the multiplexed KD, kon, koff panel back in days, not weeks.
What makes MACS® Matchmaker the purpose-built platform for degrader characterization.
20–64 Multiplexed compound capacity per chip DDI writes the entire compound library onto an 8×8 mologram array in a single pooled injection, so a 20-compound linker SAR series fits into one experiment. | r = 0.98 Method correlation across analyses Multiplexed equilibrium and kinetic KD values agree at r = 0.98, compared to 0.88 for sequential singleplex measurements on the same compound series. Within-chip conditions remove chip-to-chip drift as a SAR confounder. | Lysate-ready E3 ligases in their native context Coherent mass detection rejects non-specific binding by geometry rather than reference subtraction, so E3 ligases that are unstable purified can be presented directly from lysate. |
1:1 kinetics kon, koff, KD from one run Five-step single-cycle kinetics fits the Langmuir 1:1 model globally across all 64 spots simultaneously. Slow off-rates relevant to non-degrading molecular glues are resolved without baseline drift artifacts. | pg/mm² Direct mass readout Coherent mass density is a direct physical observable, not a refractive-index proxy. Ternary cooperativity values feed into kinetic analysis without buffer-mismatch corrections. | On-DNA hits DEL-to-lead in one workflow The DNA tag that encoded the DEL hit also drives DDI immobilization, so DEL-derived degrader scaffolds can be validated kinetically without off-DNA resynthesis. |
MACS® Matchmaker vs. competing degrader characterization platforms.
| Parameter | MACS® Matchmaker | Biacore SPR | Carterra HT-SPR | NanoTemper MST |
|---|---|---|---|---|
| Compounds per chip | ✓ 20–64 in parallel | 1–4 sequential | 384 (one ligand panel) | 1 |
| Cell lysate compatibility | ✓ Native | Limited | Limited | Quenching artifacts |
| Compound labeling | ✓ None (DNA tag is also barcode) | None | None | Fluorescent label required |
| Non-specific binding | ✓ Structurally absent | Reference subtraction | Reference subtraction | Bulk effects |
| Slow off-rate sensitivity | ✓ Drift-free baseline | Drift over long associations | Drift over long associations | Endpoint, no kinetics |
| On-DNA DEL hit validation | ✓ Native via DDI | Off-DNA resynthesis required | Off-DNA resynthesis required | Off-DNA resynthesis required |
| Setup per linker series | ✓ One pooled injection | Per-compound coupling | Spotted array | Per-compound titration |
The 20-compound DNA-VHL linker SAR study.
A peer-reviewed application note (Raschke et al., RSC Chemical Biology 2026) characterizing 20 DNA-VHL ligand conjugates with diverse amino acid linkers. Includes full kinetic and equilibrium KD data (range 20–550 nM), structure-activity correlations across cLogP, cLogS, and polar surface area, plus a side-by-side comparison of singleplex versus 20-plex multiplexed measurements.
Request the AppNote →