Validate DEL hits on-DNA, in 90 minutes.
Characterize 54 to 64 hits from a DNA-encoded library (DEL) selection in parallel on a single chip, directly from the selection eluate. No off-DNA resynthesis. No purified target. Full kinetic ranking in one run.
Why on-DNA validation closes the gap that NGS leaves open.
How DEL screening worksA DNA-encoded library (DEL) is a pool of millions to billions of small molecules, each covalently linked to a unique DNA barcode that records which building blocks were used during its combinatorial synthesis. The library is incubated with a target, non-binders are washed away, and the remaining barcodes are amplified by PCR and sequenced by next-generation sequencing (NGS). Enriched barcodes point to the chemical building blocks that produced binders. 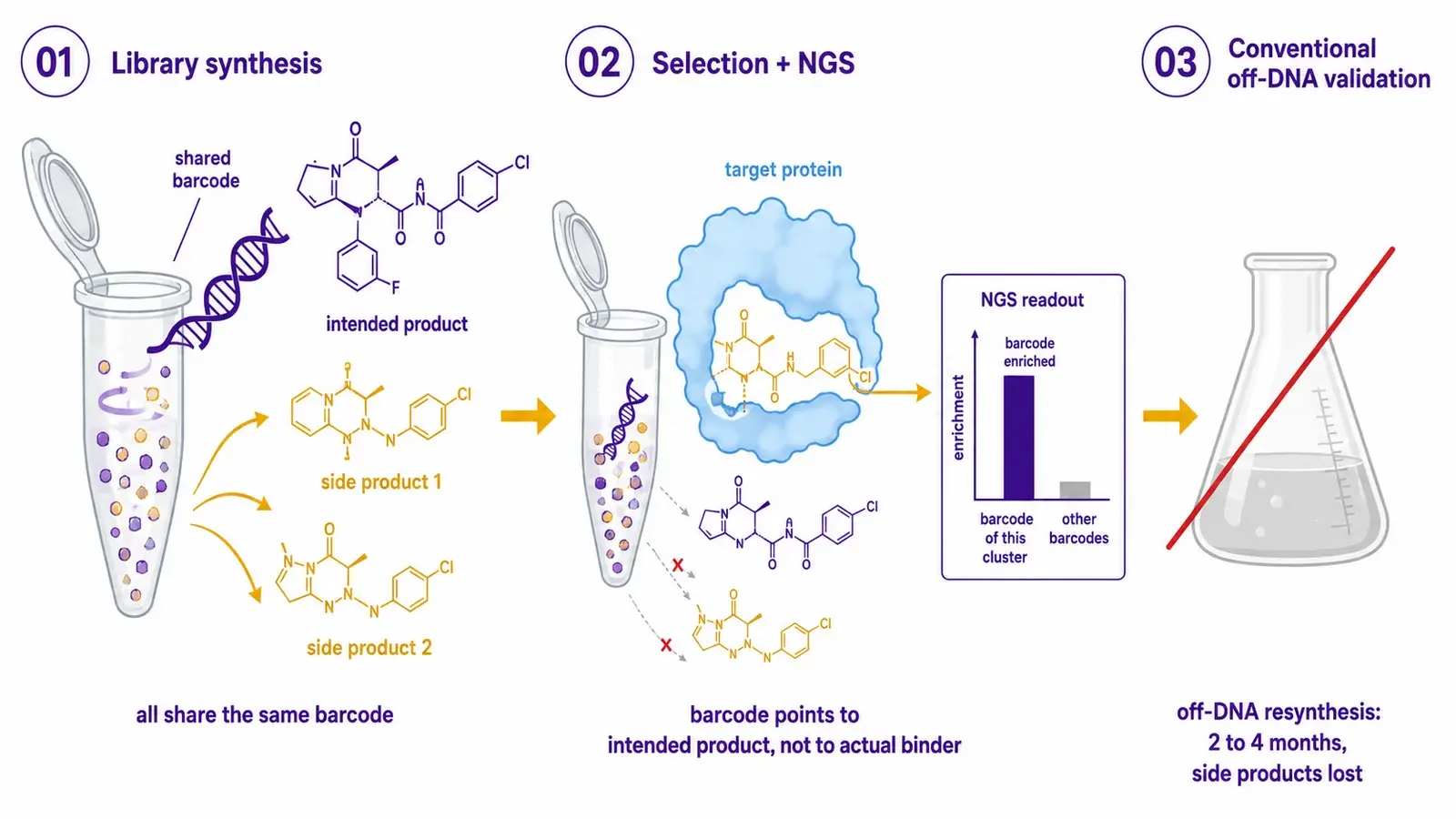 The barcode encodes the intended product, not the molecule that actually bound the target. All chemical species formed during the same synthesis step share the same barcode: intermediates, side products, on-DNA degradants, and the desired compound. Hit calling from sequencing alone cannot distinguish them. | Why off-DNA resynthesis often misses the real binderThe conventional resolution is off-DNA resynthesis: rebuild the assumed hit by traditional medicinal chemistry, then characterize. The route is fundamentally different from the original library route. Aqueous DNA-compatible chemistry is replaced by organic solvents, different stoichiometries, different reaction times. A different impurity profile is produced, and any side-product binder that drove the original enrichment is silently absent. Each resynthesis cycle takes 2 to 4 months per compound and costs a chemistry FTE per hit. Attrition rates of 60 to 80 percent are common because many resynthesized "hits" no longer bind, or bind weakly because the actual binding species was a side product that resynthesis failed to reproduce. |
One chip. 64 hits. Crude eluate. No resynthesis.
DEL hit validation demands many compounds, native DNA tags, and the freedom to measure directly in the messy output of selection. MACS® Matchmaker addresses each constraint by making DNA-directed immobilization (DDI) the native binding step, rather than an off-target afterthought.
⏱ Months of off-DNA chemistry → 90-minute panel Off-DNA resynthesis takes 2 to 4 months per hit. MACS® Matchmaker runs 54 to 64 hits in parallel on a single chip, in approximately 90 minutes, with the original on-DNA construct as the binding partner. | ∿ NGS endpoint → full kinetic profile Sequencing returns enrichment counts. MACS® Matchmaker returns k_a, k_d, and KD for each hit, so true binders, slow-off binders, and enrichment artifacts are separable on kinetic grounds, not on read count alone. |
≈ Purified target → crude selection eluate DDI captures the on-DNA hit on the chip surface; the target itself is the analyte and can be presented in selection buffer, cell lysate, or other crude matrices. No purification, no detergent screen, no immobilization chemistry per target. | ⚙ Sequence disclosure → IP-protected ligation The DNA Ligation Kit ligates a generic immobilization adapter to the proprietary DEL barcode without exposing the sequence to the chip vendor. The original barcode never leaves the customer's lab. |
Where MACS® Matchmaker accelerates DEL discovery.
The same chip and protocol address three workflows that span the full DEL discovery pipeline.
On-DNA hit validation Confirm enriched DEL hits with full kinetic characterization, directly from selection eluate, before committing medicinal chemistry resources to off-DNA resynthesis. Standard workflow at Philochem (CAIX) and the central use case at large-pharma DEL platforms. | DEL on Cells Cell-based DEL selection on cell-surface targets (G-protein-coupled receptors, ion channels, tumor antigens) produces hits that interact with native receptor topology. The MACS® Matchmaker measures these on-DNA constructs against KO-controlled cell lysate, preserving the avidity and matrix complexity of the original selection. | On-DNA SAR for adjacent modalities DNA-tagged PROTACs, peptides, glues, and ADC-DEL constructs share the on-DNA binding-validation problem. Multiplexed kinetics on the mologram array delivers structure-activity feedback before resynthesis. See the PROTAC application page for the linker-SAR workflow. |
From NGS readout to validated kinetic profile.
The 8×8 mologram array is loaded once via DDI of the on-DNA hit pool, often the selection eluate itself or a focused validation panel. The target protein is then injected in single-cycle kinetics format, and equilibrium plus kinetic KD are extracted in the same run for all spots simultaneously.
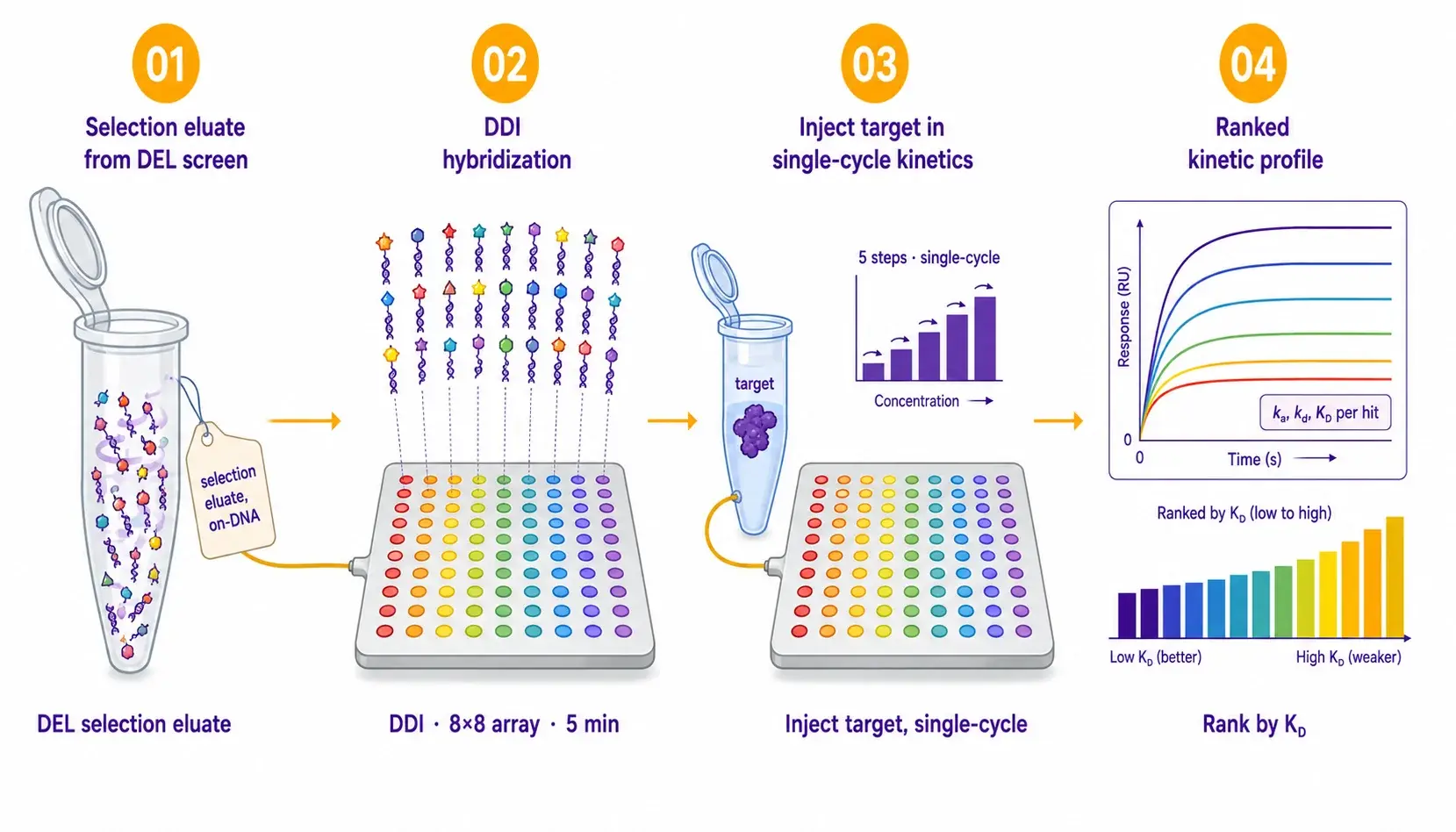 | Protocol details
|
From NGS hit to confirmed binder, in one run.
Send us your selection eluate or a focused on-DNA validation panel. Get k_a, k_d, KD for 54 to 64 hits back in days, not months.
What makes MACS® Matchmaker the purpose-built platform for DEL hit validation.
54 to 64 Multiplexed channels per chip DDI writes the entire on-DNA hit panel onto an 8×8 mologram array in a single pooled injection. Each mologram is a fully independent kinetic measurement. | <0.05 pg/mm²/min Drift-free baseline Coherent mass detection produces a baseline floor that is independent of bulk refractive-index drift. Low-affinity hits in the µM regime are resolvable in the same run as high-affinity hits in the nM regime. | Crude eluate Native sample matrix Selection eluate, cell lysate, serum, plasma, CSF, and urine are all compatible. No target purification or detergent optimization is required because non-specific binding is rejected by detection geometry. |
Label-free DNA tag is the immobilization handle The on-DNA construct is the native binding partner. No fluorescent or biotin tag is added to the compound, so DNA-tag chemistry and downstream synthetic steps remain untouched. | IP-protected DNA Ligation Kit A generic immobilization adapter is ligated to the proprietary DEL barcode in the customer lab. The barcode sequence never leaves the user's premises and is not visible to the chip or the vendor. | 1,536/day MACS® Sampler throughput The Sampler add-on automates chip loading and analyte injection so 24 chips of 64 measurements each, totaling 1,536 hit-target interactions, can be processed in a single 24-hour campaign. |
MACS® Matchmaker vs. competing DEL hit validation routes.
| Parameter | MACS® Matchmaker | Carterra HT-SPR | Biacore SPR | ASMS / Off-DNA Resynth |
|---|---|---|---|---|
| On-DNA compatibility | ✓ Native, DDI is the immobilization | Off-DNA only | Off-DNA only | Off-DNA only by definition |
| Time per validation cycle | ✓ ~90 min for 64 hits | Hours for 1 panel | Days per hit | 2 to 4 months per hit |
| Sample matrix | ✓ Crude selection eluate | Purified target only | Purified target only | Purified analyte only |
| Side-product recovery | ✓ Native (the on-DNA species is the analyte) | Lost | Lost | Lost |
| Multiplexing | ✓ 54 to 64 in parallel | 384 (one ligand panel) | 4 to 8 | 1 |
| Kinetic readout | ✓ k_a, k_d, KD per hit | k_a, k_d, KD per hit | k_a, k_d, KD per hit | Endpoint, no kinetics |
| IP / sequence disclosure | ✓ DNA Ligation Kit, sequence stays in-house | Not applicable (off-DNA) | Not applicable (off-DNA) | Not applicable |
Two peer-quality DEL Application Notes plus the published CAIX case.
The general DEL hit characterization Application Note (2024_1) introduces the on-DNA workflow, kinetic readouts, and the MACS® Matchmaker chip layout. The CAIX collaboration with Philochem (Application Note 2024_3) demonstrates the complete on-DNA validation panel for a clinically relevant tumor target. Both AppNotes include sensorgram libraries, KD distributions, and citable methods.
Request both AppNotes →