Focal Molography – A New Frontier in Biosensing
Focal molography was introduced in the latest blog article as a novel technology capable of detecting molecular interactions, without acquiring any environmental noise (e.g. temperature gradients, buffer changes, and nonspecific binding). As such, it overcomes one of the most significant hurdles label-free biosensors have faced to date, and opens the door to a new era of biosensing.
Advantages of Focal Molography
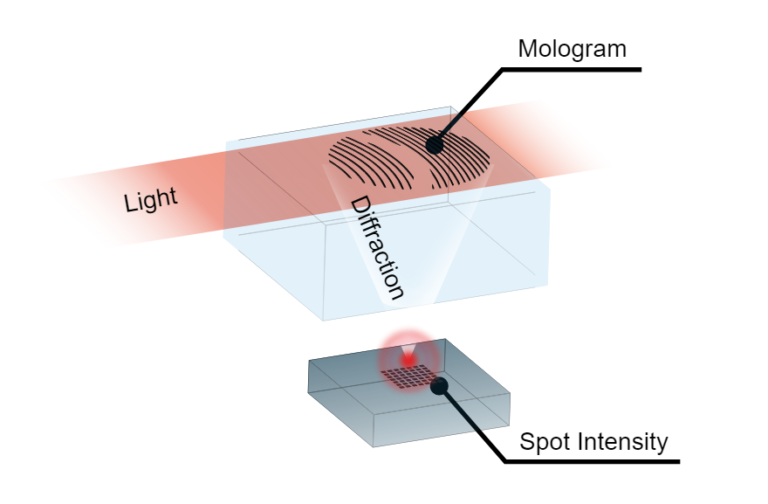
Focal molography is different than established label-free biosensors in that the molecules themselves assemble the signal generating structure. The core concept of focal molography lies in its use of a pattern of binding sites, a mologram, that has the interesting property of assembling analyte molecules such that they diffract a laser light beam into a specific point in space, the focal point. By detecting the intensity of this point, we can observe all molecular interactions happening on the grating simultaneously. Molecules with no affinity to the binding sites on the mologram (i.e. molecules we are not interested in) do not assemble on the grating; because of this, the light these molecules diffract will not be concentrated into the focal point, but rather diffracted in random directions.
This implies that interactions we do not wish to detect, i.e. environmental noise, will not be detected. With almost total rejection of environmental noise sources, focal molography brings a vast array of advantages to the table; the most important ones are discussed below.
Stable
While most refractometric biosensors such as SPR intrinsically measure and are therefore highly affected by environmental noise, focal molography remains immune to these perturbances of the signal. Indeed, as discussed in the article dedicated to SPR, refractometric biosensors detect molecular interactions based on slight changes in refractive index in the entire volume of the evanescent field. In refractometric biosensors, this volume is disproportionally large compared to the volume of the molecules present. This feature leads to the important consequence that any small change in this field (e.g. due to environmental noise) affect the sensor’s output significantly. This is not the case in focal molography: due to its nano-sized, self-referencing principle, focal molography provides unique signal stability, remaining largely unaffected by environmental influences.
Fast
In any other biosensor, signal acquisition Is affected by environmental noise and drift. It is therefore often required to reference the measurement with a secondary, parallel sensor to account for these external influences and obtain the desired readout. Focal molography is what we call “self-referencing”; this means that instead of having to compare its initial values to a known stable value from an external source from a reference channel, it can stabilize values on its own. No pre-equilibration with buffer or temperature stabilizations of the sensor are required for the measurement, which allows it to be immediately started. No matching of the refractive indices of different buffers is needed as refractive index jumps are not detected.
Robust
Focal molography is compatible with complex media, such as cell culture media and buffers, and can thereby easily be used in combination with cell-based assays. A robust technology is crucial for analysis of crude and unprocessed biological samples such as body fluids. This offers endless possibilities, as it provides a new source of important data for scientists and clinicians.
Real-Time
Biological interactions monitored in real-time (as they are happening) with focal molography. What added benefits does this bring? On a purely molecular front, real-time measurements provide information on the affinity of a molecular interaction, which can be useful information for scientists. Additionally, measuring in real-time offers insight on small and/or rapid changes occurring in the sample, which would be impossible to detect retrospectively. Finally, real-time measurements provide immediate information; this aspect is crucial in diagnostics for example, where focal molography would enable immediate readout of viral/bacterial loads in patients, but also regular screening for disease biomarkers.
Label-Free
One of the earliest blog articles discussed the advantages of label-free biosensors compared to their label-based counterparts. Briefly, label-free technologies detect the presence and/or activity of molecules of interest based on their biophysical properties, such as molecular weight, refractive index or charge, while label-based technologies require a special tag (usually another molecule), to be attached to them. The use of a label can not only alter the intrinsic properties of the molecule of interest (which can compromise the read-out), but also involves an additional and non-straightforward preparatory step. Label-free technologies are therefore of preference.
Multiplexing
Due to its label-free nature and its nano-sized principle, focal molography is very well–suited for multiplexed assays, whereby multiple assays can be run in parallel. This contributes not only to the speed of readout obtention possible, but also to amount of information that can be yielded from a single chip.
Miniaturization
Although many biosensing technologies have been developed thus far, not many have shown potential for miniaturization. Why? Surprisingly, although insufficient sensitivity is commonly assumed to be the bottleneck, this is mostly not the reason behind the impediment of their miniaturization. Here again, the limiting factor is their cross-sensitivity, in particular to environmental influences: it creates a largely disproportionate ratio of sensing volume to the size of an individual molecule. Current sensors loose sensitivity when miniaturized because they can no longer be stabilized to minimize the influence of external signal sources on the signal. However, as focal molography requires no external stabilization thanks to its self-referencing principle, a miniaturized reader will have the same performance as a large benchtop instrument, enabling miniaturization without compromising on accurate readout.
Outlook
This article sheds light on the main advantages focal molography brings to the world of biosensing. Figure 2 shows a comparison between focal molography and other current biosensing technologies, including SPR, for the attributed discussed in this article.
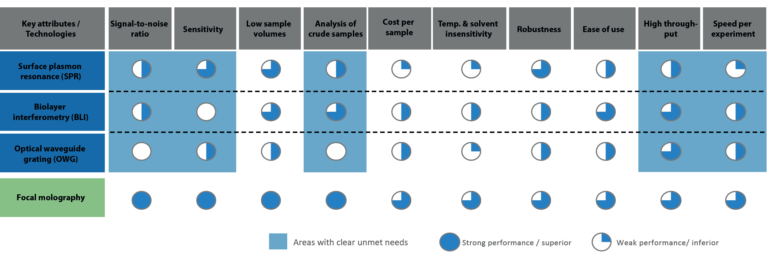
As it was hopefully demonstrated in this article, focal molography offers a novel set of possibilities for biosensing. But: What can this technology do that has not been done? How can we put its groundbreaking attributes to good use? Find out in our next article!
References
Frutiger, A., Fattinger, C. and Vörös, J., 2021. Ultra-Stable Molecular Sensors by Sub-Micron Referencing and Why They Should Be Interrogated by Optical Diffraction—Part I. The Concept of a Spatial Affinity Lock-in Amplifier. Sensors, 21(2), p.469.
Frutiger, A., Gatterdam, K., Blickenstorfer, Y., Reichmuth, A.M., Fattinger, C. and Vörös, J., 2021. Ultra Stable Molecular Sensors by Submicron Referencing and Why They Should Be Interrogated by Optical Diffraction—Part II. Experimental Demonstration. Sensors, 21(1), p.9.
2019 Kübrich Molographic Peptide Arrays Towards Label-free Protein Signaturing in Undiluted Blood Plasma

